December 9, 2020

Wheat breeders all over the world just gained a major new scientific resource.
On Nov 25, Kansas State University researchers, in collaboration with the international 10+ Wheat Genomes Project led by the University of Saskatchewan, announced the complete genome sequencing of 15 wheat varieties representing breeding programs around the world — an invaluable resource to improve global wheat production.
This effort gained momentum in 2018, when the Kansas State University team, in collaboration with the International Wheat Genome Sequencing Consortium, published the genome assembly of the wheat variety Chinese Spring, the first complete reference genome of bread wheat. With rapid advances in DNA sequencing technology, and with experience from assembling the first wheat genome, the 10+ Wheat Genomes Project brought together the expertise and resources of more than 95 scientists from universities and institutes in Canada, Switzerland, Germany, Japan, the United Kingdom, Saudi Arabia, Mexico, Israel, Australia and the United States.
This study represents the start of a larger effort to generate thousands of genome sequences of wheat, including genetic material brought in from wheat's wild relatives.
“It’s like finding the missing pieces for your favorite puzzle that you have been working on for decades,” says Curtis Pozniak, the project leader, a wheat breeder and the director of the University of Saskatchewan Crop Development Centre. “By having many complete gene assemblies available, we can now help solve the huge puzzle that is the massive wheat pan-genome and usher in a new era for wheat discovery and breeding.”
K-State well-suited to study
Jesse Poland, associate professor at K-State and director of the Feed the Future Innovation Lab for Applied Wheat Genomics and the Wheat Genetics Resource Center-University Cooperative Research Center, says the K-State team was uniquely suited to represent U.S. wheat in the effort.
“We are fortunate to have world leaders in breeding and genetics under one roof, and generous support from the National Science Foundation, Kansas Wheat and many others," Poland says.
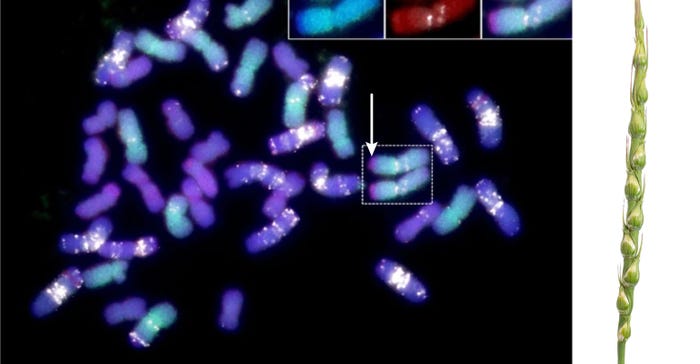 WHEAT CHROMOSONE: At left, a chromosome segment from Aegilops ventricosa, highlighted by the white arrow, is found in the Kansas wheat variety Jagger. This segment carries important genes for pest and disease resistance, and is associated with increasing grain yield. (Courtesy of Dal-Hoe Koo) The image on the right is this plant, wild wheat relative Aegilops ventricosa. (Courtesy of Haley Ahlers)
WHEAT CHROMOSONE: At left, a chromosome segment from Aegilops ventricosa, highlighted by the white arrow, is found in the Kansas wheat variety Jagger. This segment carries important genes for pest and disease resistance, and is associated with increasing grain yield. (Courtesy of Dal-Hoe Koo) The image on the right is this plant, wild wheat relative Aegilops ventricosa. (Courtesy of Haley Ahlers)

The Kansas team was responsible for sequencing and analyzing the hard red winter wheat variety Jagger, released in 1994 by the Kansas State University breeding program, now led by Allan Fritz. Jagger was a landmark wheat variety in the Great Plains and covered millions of acres for many years. It was selected for this project because of its relevance as a breeding parent, as it is found in the pedigrees of current varieties across the U.S.
“Because of our collaboration in this project, we’ve had access to this phenomenal genomics resource as it’s been built, which has already led to tremendous discovery,” Poland says. “K-State plant genetics graduate student Emily Delorean is using data from the 10+ Wheat Genomes Project to develop a comprehensive analysis of important quality genes and develop better molecular breeding tools, which will have a huge impact on bread making.”
In a companion piece published in Theoretical and Applied Genetics, K-State scientists Liangliang Gao, Dal-Hoe Koo and their team completed detailed characterization of the 2N introgression, a chromosome segment that was transferred from wild wheat relative Aegilops ventricosa, which is found in Jagger but was not present in the original Chinese Spring reference genome. The 2N segment possesses resistance genes to multiple wheat diseases, including stem and leaf rust, nematodes and the emerging wheat blast disease. The team found that this chromosome segment is present in about 80% of Kansas wheat lines and also a large proportion of wheat around the world, marking its importance toward addressing global wheat improvement.
“Progress of this magnitude is only possible because of the strength of the international wheat breeding network and strong international collaborations in wheat research,” says Justin Gilpin of Kansas Wheat. “It is exciting for the Kansas wheat growers to be part of this excellent work.”
The work at Kansas State University was supported by the National Science Foundation, Kansas Wheat, the U.S. Agency for International Development, and the National Institute of Food and Agriculture. Learn more at 10wheatgenomes.com. See a complete list of international funding partners available at 10wheatgenomes.com/funders.
Source: Kansas State University News Service, which is solely responsible for the information provided and is wholly owned by the source. Informa Business Media and all its subsidiaries are not responsible for any of the content contained in this information asset.
You May Also Like




