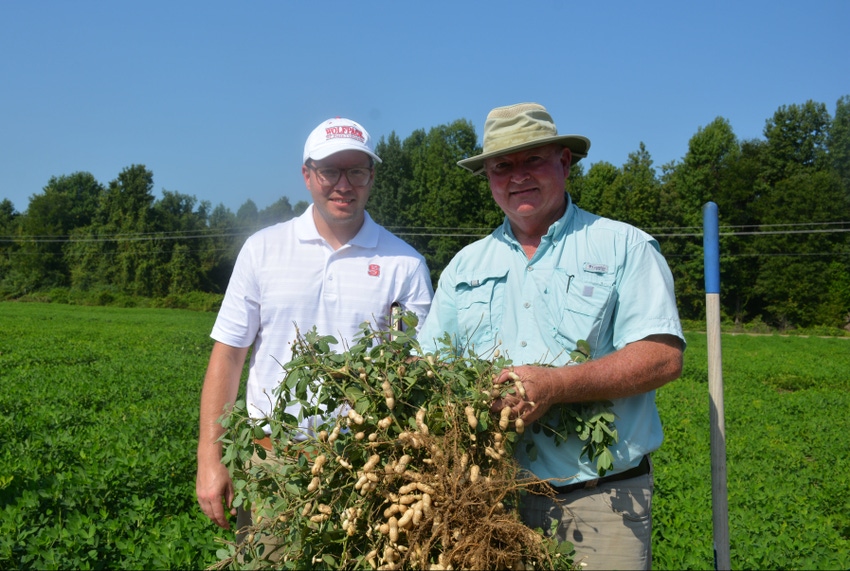
Like his predecessor, Dr. Jeff Dunne, North Carolina State University’s new peanut breeder, studied at Michigan State University. He’s hoping it’s a good sign the successful breeding work of Dr. Tom Isleib, North Carolina State’s longtime peanut breeder, will continue.
“I went to Michigan State like Tom, so we might have some inherent ability to breed peanuts at North Carolina State,” Dunne said at the peanut field day Sept. 6 at the Peanut Belt Research Station in Lewiston-Woodville. It was Dunne’s first field day speaking as North Carolina State’s peanut breeder.
Isleib retired this spring, moving to Michigan with his wife Sandy.
Dunne mentored under Isleib earlier this year prior to the retirement. Isleib began his career as North Carolina State’s peanut breeder in 1990, serving 28 years in the role. He is the father of the highly popular disease-resistant Virginia cultivars Bailey and Sugg.
Dunne earned his Bachelor’s and Master’s degree in turfgrass management from Michigan State while Isleib also earned his undergraduate degree in crop and soil science there. Both Dunne and Isleib earned their Ph. Ds. from North Carolina State. Dunne’s Ph.D. is also in turf grass management.
At North Carolina State, Dunne worked with Dr. Susana Milla-Lewis in creating a new bermudagrass breeding program. His Ph.D. worked focused on shade and freezing tolerance of bermudagrass.
He also worked in Dr. Jim Holland’s maize breeding and genetics program with USDA’s Agricultural Research Service in Raleigh prior to becoming North Carolina State’s peanut breeder.
Dunne lettered in hockey as an undergraduate at Michigan State. “No, I don’t have all my teeth. These are kind of fake,” Dunne said with a smile at the Lewiston-Woodville field day.
Like Isleib, Dunne said his peanut breeding work will focus on disease resistance and improving yield. However, a new focus will be implementation of marker assisted selection and genomic selection to develop improved peanut cultivars.
At the peanut field day in Lewiston-Woodville, Dunne explained how both marker assisted selection and genomic selection can work.
“I relate marker assisted selection to a highway road sign,” Dunne said.
“There are 20 highways in the peanut genome. Each one of these highways have different exits, and at each one of these exits we have a number of signs or markers that tell you whether there is food available or lodging available,” he said.
“Not all of these are consistent across all the exits, so what we want to be able to do is find those exits across those 20 highways that are within the peanut genome and be able to select on those because there is some kind of association between this exit and pod color or pod characteristics of the high-oleic trait.”
In his work, Dunne plans to implement marker assisted selection by focusing on a few of the easier selected traits such as pod characteristics or the high oleic trait. “We will select on those until we start developing our populations or our breeding lines that have a large component of those markers that are present,” Dunne said.
Genomic selection will also be a key part of Dunne’s breeding work. Using the same metaphor of 20 highways in the peanut genome, he says it’s important to discover how long it takes to travel from one highway to another.
“You plug this address into Google maps, and it instantly spits out some sort of estimate as how long your trip will take based on construction or traffic patterns, whether you picked the fastest route or whether you picked the one with the lowest number of miles,” he explains.
“All of these factors are going into estimating how long it will take you to get to your destination. In terms of genomic selection, we’re doing kind of the same thing. We’re using these same very markers we did for marker assisted selection, and we are going to try to make estimates on how good the yield is prior to even putting it in the field. This allows us to save time and increase our efficiency. We are essentially only evaluating the things we really want to have in the field.”
Dunne is settling in as North Carolina State’s new peanut breeder. He has been making the rounds, speaking at the 50th annual meeting of the American Peanut Research and Education Society (APRES) in Williamsburg, Va., in July and was on hand to answer questions at the Severn Field Day at the John Tyndall Farm in Pikeville in September.
Dunne has hired a research technician, Andrew Oakley to help him in his work.
Dr. Ryan Andres has been hired as research associate and will head the molecular breeding lab for the peanut breeding and genetics program at North Carolina State.
Dunne says his team is now in place to develop exceptional peanut varieties.
About the Author(s)
You May Also Like